MedComm-Oncology | Clinical homologous recombination repair gene reversion analysis identifies mechanisms of resistance to PARP inhibitors and platinum-chemotherapy

Open the phone and scan
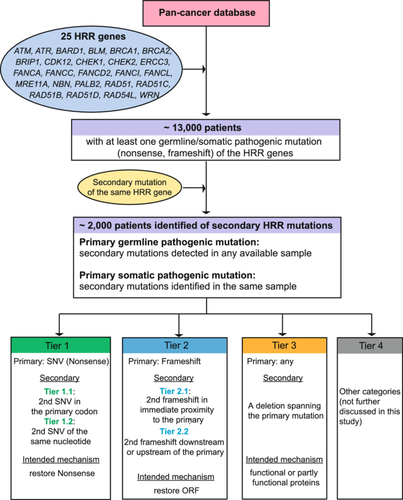
Study design. The 25 pre-defined HRR genes covered by the NGS panel is shown. Patient screening flow is demonstrated with the rationale of the four-tier classification. HRR, homologous recombination repair; NGS, next-generation sequencing; SNV, single nucleotide variant.
Identifying mechanisms underlying cancer resistance to therapy is vital for advancing treatment strategies. Pathogenic mutations of homologous recombination repair (HRR) genes are known biomarkers for platinum (Pt)-based chemotherapy and poly ADP ribose polymerase inhibitors (PARPi) effectiveness. Yet, the dynamics of HRR reversion mutations, which may herald therapy resistance, are not fully elucidated. Addressing this gap, our study analyzed secondary HRR gene mutations in a comprehensive pan-cancer data set of approximately 13,000 patients who underwent targeted next-generation sequencing. We identified a subset of patients harboring secondary mutations, which were further categorized into three tiers based on their nature, and occur in the presence of a primary pathogenic mutation, notably in BRCA1, BRCA2, PALB2, and RAD51D genes. Here we show that secondary BRCA2 mutations, indicative of adaptive resistance, emerge post-Pt/Olaparib treatment. This challenges the prevailing notion that pathogenic HRR mutations uniformly predict therapeutic sensitivity, highlighting a nuanced genetic interplay that impacts treatment success. This investigation enriches our understanding of cancer's adaptive mechanisms against therapy, suggesting a pivotal shift towards more personalized, dynamic treatment strategies. It underscores the imperative of adapting to cancer's genetic evolution, aiming for a step ahead in the ongoing battle against this disease.
Article Access: https://doi.org/10.1002/mog2.79
More about MedComm-Oncology: https://onlinelibrary.wiley.com/journal/27696448
Looking forward to your contributions.


