MedComm | Insights on the structure–function relationship of human multidrug resistance protein 7 (MRP7/ABCC10) from molecular dynamics simulations and docking studies

Open the phone and scan
ATP-binding cassette (ABC) transporters superfamily mediates multidrug resistance in cancer by extruding structurally distinct chemotherapeutic agents, causing failure in chemotherapy. Among the 49 ABC transporters, multidrug resistance protein 7 (MRP7 or ABCC10) is relatively new and has been identified as the efflux pump of multiple anticancer agents including Vinca alkaloids and taxanes. Herein, we construct and validate a homology model for human MRP7 based on the cryo-EM structures of MRP1. Structure–function relationship of MRP7 was obtained from molecular dynamics simulations and docking studies and was in accordance with previous studies of ABC transporters. The motion patterns correlated with efflux mechanism were discussed. Additionally, predicted substrate- and modulator-binding sites of MRP7 were described for the first time, which provided rational insights in understanding the drug binding and functional regulation in MRP7. The findings will benefit the high-throughput virtual screening and development of MRP7 modulators in the future.

Clinically, MRP7 has been reported to play important roles in acquired MDR and the prognosis of certain cancers. Additionally, MRP7 participates in FOXM-induced 5-FU resistance in colorectal cancer patients based on the strong correlation between mRNA levels of MRP7 and FOXM in tumor tissues. Furthermore, MRP7 also contributes to alteration in intracellular permeation of nevirapine, a nonnucleoside reverse transcriptase inhibitor for HIV-1. As a result, further understanding the structural features and transportation mechanisms of MRP7 is crucial in increasing the survival rate of patients with limited response to chemotherapy due to acquired MDR, as well as developing and discovering substrates/modulators to overcome MRP7-mediated MDR or decreasing unexpected synergistic toxicity in combinational chemotherapies. However, due to the difficulty and cost in obtaining crystal structure of membrane protein, no high-resolution structure of human MRP7 is available so far. Previously, the cryo-EM structure of bovine MRP1 was reported at 3.1–3.5 Å with inward- and outward-facing conformations. Although there are significant differences between MRP1 and MRP7 in length, size, amino acid sequence, and transportation pattern, it still provides possibility to construct high-quality homology models of MRP7 via computational strategies. Here, we present the homology models of MRP7 combining current knowledge of the transporter and the homology modeling methods based on the cryo-EM structure of MRP1 in order to (1) provide a functionally validated human MRP7 homology model; (2) assess the structural dynamics to identify potential conformational changes associated with efflux mechanism; and (3) evaluate the behavior of reported substrates and modulators of MRP7 by analyzing ligand–protein interactions. Also, extra docking simulations were performed using template MRP1 and our homology model to further validate that the MRP7 model was not biased toward its templates. The study will provide rational insights in the drug development or repurposing of MRP7 modulators.
The final model is shown in Fig. 1. In this study, TMD0 was not included in our homology model due to the fact that the TMD0s in the templates were not completely resolved. Furthermore, previous studies have shown that removal of TMD0 did not affect the function of MRP1 protein. According to the sequence alignment, MRP1 and MRP7 share similar structural domains including TMDs, Lasso (L0 linker), and NBDs (Figure 1B–1F). The alignment contains conserved region of NBDs shared by multiple ABCC members including the Walker A, Signature, and Walker B6. The consensus ATPase sites are used for establishing the Mg-ATP system.
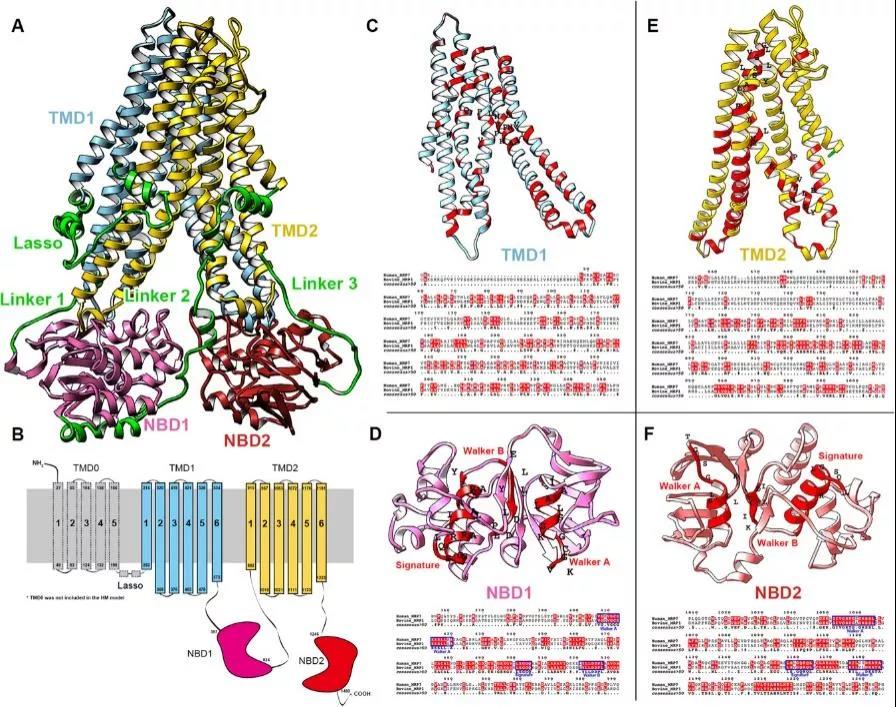
Fig. 1 Sequence alignment of MRP1 and MRP7. (A) Homology model of human MRP7
Article Access: https://onlinelibrary.wiley.com/doi/10.1002/mco2.65
Looking forward to your contributions.


